Researchers used synthesized DNA to store and read data
 EMBL-European Bioinformatics Institute (EMBL-EBI) researchers have developed a method to store and read information from deoxyribonucleic acid (DNA) molecules. The researchers devised a method to convert data into DNA code, and used it to encode 739 kilobytes of data into actual DNA. The information was retrieved with 100% accuracy by sequencing the sample and reconstructing the original files.
EMBL-European Bioinformatics Institute (EMBL-EBI) researchers have developed a method to store and read information from deoxyribonucleic acid (DNA) molecules. The researchers devised a method to convert data into DNA code, and used it to encode 739 kilobytes of data into actual DNA. The information was retrieved with 100% accuracy by sequencing the sample and reconstructing the original files.
An estimated worth of digital information in the world is about three zettabytes (3 billion terabytes) and the volume of data is constantly growing. Conventional technologies for digital data storage such as hard disks, solid state drives, and magnetic tape have their limits. Hard disks need constant supply of electricity and are expensive. On the other hand, magnetic tape doesn’t require any power to storage, but they degrade and must be replaced every ten years.
By contrast, DNA molecule is stable, durable, tense and doesn’t require any power for digital information storage. DNA-based storage of digital data could theoretically store about 100 million hours of high-definition video in about a cup of DNA.
“We already know that DNA is a robust way to store information because we can extract it from bones of woolly mammoths, which date back tens of thousands of years, and make sense of it”, said Nick Goldman, a computational biologist at the European Bioinformatics Institute in Hinxton, England.
There have been many obstacles in realization of the actual DNA–based storage system. Reading from DNA and writing to it are prone to errors which occur when the same DNA letter is repeated. Another problem is the length of data since current technology allows scientists to create only short strands of DNA. However, new method dramatically reduces the copying errors and overcomes size problems.
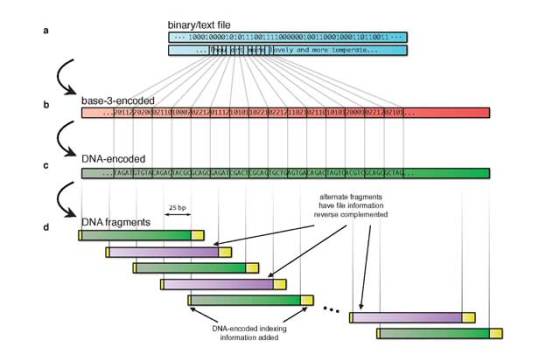
The new method for storage information involves breaking up the files into thousands of overlapping fragments, with indexing instruction for restoration of the fragments in the proper order when the data is being read. Also, the new coding scheme devised by the EMBL-EBI researchers reduces the possibility of repeating letters. The code would fail if the same error occurred on four different fragments, which would be extremely rare.
In cooperation with Agilent Technologies, Inc, a California-based company, the researchers have synthesized hundreds of thousands of pieces of DNA from a number of encoded files in order to test their technique. The encoded file were an audio recording of the Martin Luther King Jr. “I Have A Dream” speech, ”; a .jpg photo of EMBL-EBI; a .pdf of Watson and Crick’s seminal paper, “Molecular structure of nucleic acids” and a .txt file of all of Shakespeare’s sonnets.
“We’ve created a code that’s error tolerant using a molecular form we know will last in the right conditions for 10,000 years, or possibly longer”, said Nick Goldman. “As long as someone knows what the code is, you will be able to read it back if you have a machine that can read DNA.”
DNA could remain readable for such a long time as long as it is kept somewhere cool, dark and dry. However, there are some downsides to DNA as a data-storage medium. The researchers needed two weeks to reconstruct their five files, although they claim it could be done within a day with a better equipment or by adding more sequencing machines.
For more information, read the article published in Nature: “Towards practical, high-capacity, low-maintenance information storage in synthesized DNA”.









I would have to add more downsides, such as the time needed to read the data as well as its durability when it comes to neutron bombs or their equivalents.
Another huge disadvantage is its current price. According to researchers, storing 1MB of data costs around $12,400, and they claim some improvements could lead to a price of $7,440/MB. But hey, 5MB of data storage sold by Apple back in 1980 was $3,500.
Nevertheless, this is one of the most important recent breakthroughs, and it could lead to various new technologies that could be used to store and read data in synthesized DNA.